Genetic Variants Identified In TNF Gene of Patients with Juvenile Systemic Lupus Erythematosus
Written by |
In a new study entitled “Tumor necrosis factor-alpha single nucleotide polymorphisms in juvenile systemic lupus erythematosus” researchers investigated genetic variants, specifically those known as single nucleotide polymorphisms, within the gene tumor necrosis factor-alpha (TNF-a), a potent pro-inflammatory cytokine that was previously suggested to play a role in juvenile systemic lupus erythematosus (JSLE). The team found variations in the TNF-a gene that may influence patients’ susceptibility to JSLE. The study was published in the journal Human Immunology.
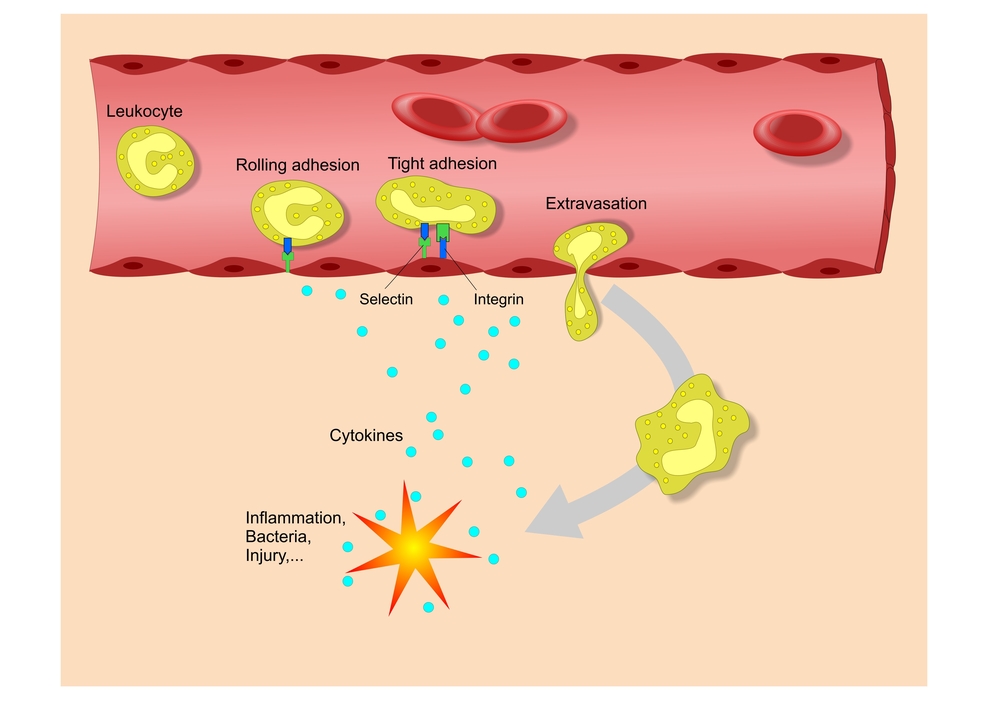
Juvenile systemic lupus erythematosus (JSLE) is an autoimmune disease that manifests itself in patients under 16 years old and is characterized by inflammatory damages, affecting multiple systems including joints, kidney, central nervous system, and hematopoietic system (bone marrow and lymph nodes). A crucial player identified in the pathogenesis of JSLE is the deregulated production of cytokines (signaling molecules that work as a communication vehicle in immune responses and stimulate migration of immune cells towards inflammatory sites) that prompt inflammation, such as tumor necrosis factor-alpha (TNF-a).
In the study, authors investigated the presence of genetic variants (specifically, Single Nucleotide Polymorphisms, SNP) within the TNF-a gene in a subset of juveline SLE patients and how they associate with the immunopathology of the disease. To this end, the team analyzed the TNFA gene in blood samples retrieved from fifty-nine patients with JSLE (10 male, 49 female) and healthy controls (137). Patients with JSLE were enrolled from the Rheumatology Clinic of the Children’s Medical Center Hospital, the Pediatrics Center of Excellence in Iran.
They found that a particular SNP (occurring at position -238 in the TNFa gene) exhibited a significantly higher frequency in JSLE patients when compared to healthy controls; additionally, the team discovered other variations within the TNFA gene occurring specifically in JSLE patients.
Thus, while in this study authors established the existence of SNPs in JSLE patients, they do recognize limitations to their study, including the relatively small number of participants and the fact that they did no correlation between the SNPs identified in JSLE patients and the clinical manifestations of the disease. However, the team suggests the identified genetic variants in larger sample size studies could identify new genetic risk factors associated with the development of JSLE.

